Before we begin, I should disclose: I am not a believer in the semantic web. I am not excited by the promise of linked data.
As I mentioned when I was getting started with SPARQL a few posts ago, I’ve been convinced to give this a try because Cray, Inc. has a new type of computer and this may be the lowest-overhead way for me to use it for global health metrics.
With that out of the way, here is one way that I may test drive their machine: counting triangles in massive graphs. I’ll abbreviate the introduction, counting triangles is an area that there has been a fair amount of work on in the last decade. Google scholar can get you more up-to-date than I, although I was looking into this matter towards the end of my post-doc at Microsoft Research. It is a good simplification of a more general subgraph counting challenge, and it can probably be justified in its own right as a metric of “cohesion” in social networks.
Another appealing aspect of triangle counting is that it is easily done with the Python NetworkX package:
import networkx as nx G = nx.random_graphs.barabasi_albert_graph(10000, 5) print 'Top ten triangles per vertex:', print sorted(nx.triangles(G).values(), reverse=True)[:10]
It is not as easy, but also not much harder to count triangles per vertex in SPARQL (once you figure out how to transfer a graph from Python to a SPARQL server):
SELECT ?s (COUNT(?o) as ?hist)
WHERE { ?s ?p ?o .
?o ?p ?oo .
?s ?p ?oo .
}
GROUP BY ?s
ORDER BY DESC(?hist) LIMIT 10
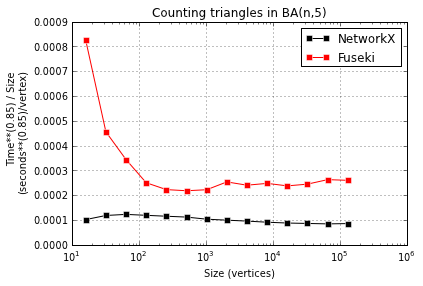
I compared the speed of these approaches for a range of graph sizes, but just using the Jena Fuseki server for the SPARQL queries. Presumably, the Cray uRiKa will be much faster. I look forward to finding out!
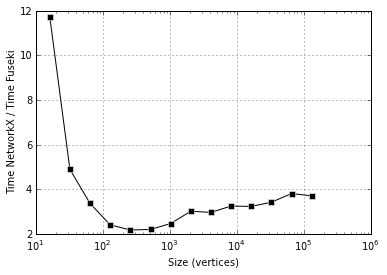
NetworkX is faster than Fuseki, 2-4x faster. But more important is the next plot, showing that both seem to take time super-linear in instance size, possibly with different exponents: