This post is a little tutorial on how to use PyMC to sample points uniformly at random from a convex body. This computational challenge says: if you have a magic box which will tell you yes/no when you ask, “Is this point (in n-dimensions) in the convex set S”, can you come up with a random point which is nearly uniformly distributed over S?
MCMC has been the main approach to solving this problem, and it has been a great success for the polynomial-time dogma, starting with the work of Dyer, Frieze, and Kannan which established the running-time upper bound of . The idea is this: you start with some point in S and try to move to a new, nearby point randomly. “Randomly how?”, you wonder. That is the art.
In PyMC, if you want to sample points uniformly from the unit square, all you need to do is define your stochastic variable, make a Markov chain, and sample from it. Here is how:
from pymc import *
X = Uniform('X', zeros(2), ones(2))
mc = MCMC([X])
mc.sample(10000)
If you have matplotlib set up right, you should be able to get a nice picture of this now
plot(X.trace()[:,0], X.trace()[:,1],',')
Not a very complicated convex body, but it’s a start. And it was easy! Five lines of code? No problem.
Say you want something to sample from some more complicated, though, like anything-that’s-not-a-rectangle. One way to do this is to use PyMC’s potential object. Below is a constraint that makes S into a ball of diameter one, centered at (0.5, 0.5). It just says that the log-probability for any point outside of this ball is
from pymc import *
X = Uniform('X', zeros(2), ones(2))
@potential
def s1(X=X):
if ((X-0.5)**2).sum() >= 0.25:
return -inf
else:
return 0.0
Did that give you an error? For a fraction of readers, it will; PyMC gets unhappy when it has a sample with probability zero. Try setting X to something in the ball first:
from pymc import *
X = Uniform('X', zeros(2), ones(2))
X.value = [0.5,0.5]
@potential
def s2(X=X):
if ((X-0.5)**2).sum() >= 0.25:
return -inf
else:
return 0.0
mc = MCMC([X, s2])
mc.sample(10000)
plot(X.trace()[:,0], X.trace()[:,1],',')
But, this is too easy to sample from. It is a nice, plump convex body. The hard stuff comes from the convex bodies with skinny bits. If you live in high dimensions, these skinny bits show up everywhere. For example, if you have a ball just like the one above, but in 20 dimensions instead of 2, check out what happens:
from pymc import *
X = Uniform('X', zeros(20), ones(20), value=0.5*ones(20))
@potential
def s3(X=X):
if ((X-0.5)**2).sum() >= 0.25:
return -inf
else:
return 0.0
mc = MCMC([X, s3])
mc.sample(10000)
plot(X.trace()[:,0], X.trace()[:,1],',')
That’s not looking nice anymore. The problem is this. If you don’t tell PyMC how you want the Markov transitions to go, it picks something for you. It defaults to a Metropolis walk, where each stochastic variable is drawn from its prior distribution. In this case, that means that for 10,000 samples, PyMC draws uniformly from the 20-dimensional hypercube, and then rejects it and stays at
if
falls outside of the ball we want. Since the volume of the ball is much smaller than the volume of the cube, roughly 75% of these samples are rejected. (In the picture above, I jittered the points to reveal the duplicate points.)
How can we make things nice again? Instead of sampling from the prior every time, we can take a small, random step away from the point that we are currently at.
from pymc import *
X = Uniform('X', zeros(20), ones(20), value=0.5*ones(20))
@potential
def s4(X=X):
if sum((X-0.5)**2) >= 0.25:
return -inf
else:
return 0.0
mc = MCMC([X, s4])
mc.use_step_method(Metropolis, X, dist='Normal', sig=0.05)
mc.sample(50*10000,thin=50)
plot(X.trace()[:,0], X.trace()[:,1],',')
The new trick is in line 10, where I asked for a particular step method. It appears that the dist parameter tells PyMC to use move to a new point normally distributed around the current point, and the sig parameter controls the variance of the normal distribution. Now things looks nice again. Maybe this isn’t actually a great example, because it takes longer for this chain to mix, so the pictures that look nice come from keeping 1 sample out of every 50, meaning the chain takes 50 times longer to run. If I really wanted samples from the 20-dimensional ball, rejection sampling is a better way to go. (But if I really wanted samples from the 20-dimensional ball, there are even better ways to draw them…)
Up next, how to implement the hit-and-run walk in PyMC. (Actually, not next…)




Abie,
What particular rejection sampling algorithm do you have in mind in the second-to-last sentence?
Anand
Anand: Let me see if I am learning the language of stats MCMC. I think that you would say that the hit-and-run walk uses a Gibbs method, because it never rejects a sample.
The way it works (in the paper by Lovasz and Vempala linked above, when specialized to uniform sampling) is to: pick a uniformly random line through the current point, and move to a uniformly random point on the line and inside the set.
This is the walk which currently has the best upper bound on mixing time. From a “warm-start” it requires steps, a huge improvement on the original proof that a poly-time procedure exists, which had the
steps, a huge improvement on the original proof that a poly-time procedure exists, which had the 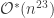 bound I mentioned above.
bound I mentioned above.
I think the name is taken from sports (not traffic-related injuries).
Huh, that’s really cool. I’ll have to check out the paper. Do you know if it can be mixed into a larger MCMC algorithm, ie if some variables are being updated with hit-and-run and some are being updated with Metropolis, do you still get joint samples?
Gibbs methods update variables by drawing them conditional on their Markov blanket. It doesn’t seem like hit-and-run is doing that, so you probably wouldn’t call it a Gibbs method even though it never rejects a jump.
Incidentally, the default step method being used for X is the one in the ref below. Rather than proposing from the prior, it proposes from a multivariate normal centered on the current state. The covariance is a scalar multiple of its estimate of the covariance of the ‘posterior’, which it updates on the fly.
Heikki Haario, Eero Saksman, and Johanna Tamminen. An adaptive Metropolis algorithm. Bernoulli, 7(2):
223–242, 2001.
Ananda: It seems like it should be possible. I think that it is straight-forward to show that it works for uniform sampling from a convex body you can mix hit-and-run on some dimensions and another method on the other dimensions.
Where things get more complicated would be when you are approximating hit-and-run in a space that is not uniform (and probably not even log-concave). My guess is that the stationary distribution of the chain will be within of the true stationary distribution.
of the true stationary distribution.
Of course, the question remains, is this actually a good idea in practice.
I’ll look at the Haario, et al paper in more detail soon. It reminds me of a different theoretical result by Vempala. It does mean that the default behavior of PyMC is non-Markovian, right? 🙂
We hang our heads in shame. 🙂 See Levine et al. Implementing componentwise Hastings algorithms. Computational Statistics & Data Analysis (2005) vol. 48 (2) pp. 363-389
We await part 2! I want to see the code for a modern convex sampler…
Thanks for the encouragement… this hasn’t even made it on my to-do list yet, but I would love to come back to it one of these days.
I consider PyMC a modern convex sampler! Actually, I should re-write some portions of this tutorial to more accurately reflect what PyMC 2.0 (just released, congratulations!) is doing.
And the great thing about PyMC is that it’s open source and written in python, so you can see the code with one click.
To Read: arxiv paper on when the Adaptive Metropolis will still converge to the intended distribution.
Eero Saksman, Matti Vihola, On the Ergodicity of the Adaptive Metropolis Algorithm on Unbounded Domains.
Two years ago, I was just joking around about writing a hit-and-run sampler, but I finally got off my butt and did a proof of concept! Code and movie at the end of this post: https://healthyalgorithms.wordpress.com/2011/01/28/mcmc-in-python-pymc-step-methods-and-their-pitfalls/