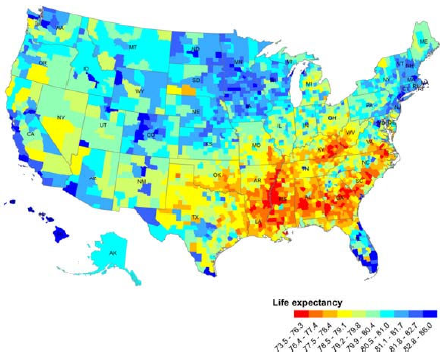
This recent study by my colleagues has been making headlines a lot last week, but I’m just getting to write about it now. While I was busy, stories about it appeared in high-profile outlets like NPR and the Statistical Modeling, Causal Inference, and Social Science blog.
As I’ve been thinking for two years (according to the ancient post I pushed out the door yesterday), life expectancy is a weird statistic. Life expectancy at birth is not, as the name might imply, a prediction on the average length of the life of a baby born this year. It is something more complicated to describe, but easier to predict. I like to think of it as the length of life if you froze the world exactly the way it is right now, and the baby today was exposed to the mortality risk of today’s one-year-olds next year, today’s two-year-olds in two years, etc. Although, as a friend pointed out two weeks ago, this is not a really good way to look at things either, if you push the analogy too hard. Currently Wikipedia isn’t really helpful on this matter, but maybe it will be better in the future.
There is another interesting thing in this paper, which is the validation approach the authors used. Unfortunately, it’s full development is in a paper still in press. Here is what they have to say about it so far:
We validated the performance of the model by creating small counties whose “true” underlying death rates were known. We did this by treating counties with large populations (> 750,000) as those where death rates have little sampling uncertainty. We then repeatedly sampled residents and deaths from these counties (by year and sex) to construct simulated small-county populations. We used the above model to predict mortality for these small, sampled-down counties, which were then compared with the mortality of the original large county.
I believe that this is fully developed in the paper which they cite at the beginning of the modeling section, Srebotnjak T, Mokdad AH, Murray CJL: A novel framework for validating and applying standardized small area measurement strategies, submitted. From what I’ve heard about it, I like it.