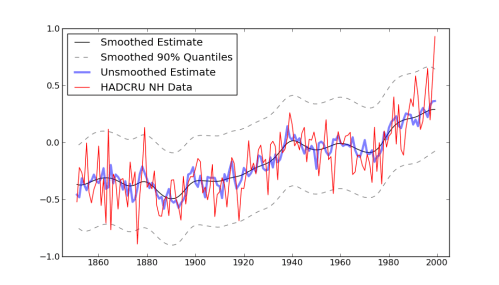
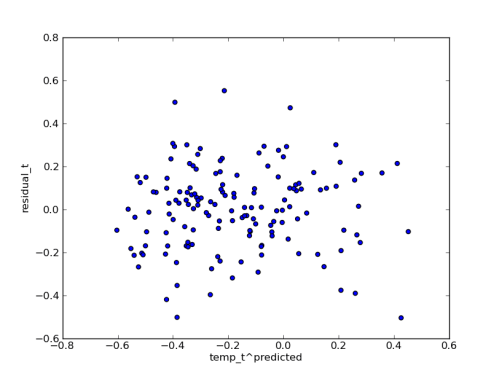
I haven’t had time to say much recently, due to some travel for work, but I did have a chance to prototype the Hit-and-Run/Adaptive Metropolis approach to MCMC that I mentioned in my last post. Was that really three weeks ago? How time flies.
Anyway, thanks to the tip Aram pointed out in the comments, Hit-and-Run can take steps in the correct direction without explicitly computing an approximation of the covariance matrix, just by taking a randomly weighted sum of random points. It’s nearly magic, although Aram says that it actually makes plenty of sense. The code for this “H-RAM” looks pretty similar to my original Hit-and-Run step method, and it’s short enough that I’ll just show it to you:
class HRAM(Gibbs):
def __init__(self, stochastic, proposal_sd=None, verbose=None):
Metropolis.__init__(self, stochastic, proposal_sd=proposal_sd,
verbose=verbose, tally=False)
self.proposal_tau = self.proposal_sd**-2.
self.n = 0
self.N = 11
self.value = rnormal(self.stochastic.value, self.proposal_tau, size=tuple([self.N] + list(self.stochastic.value.shape)))
def step(self):
x0 = self.value[self.n]
u = rnormal(zeros(self.N), 1.)
dx = dot(u, self.value)
self.stochastic.value = x0
logp = [self.logp_plus_loglike]
x_prime = [x0]
for direction in [-1, 1]:
for i in xrange(25):
delta = direction*exp(.1*i)*dx
try:
self.stochastic.value = x0 + delta
logp.append(self.logp_plus_loglike)
x_prime.append(x0 + delta)
except ZeroProbability:
self.stochastic.value = x0
i = rcategorical(exp(array(logp) - flib.logsum(logp)))
self.value[self.n] = x_prime[i]
self.stochastic.value = x_prime[i]
if i == 0:
self.rejected += 1
else:
self.accepted += 1
self.n += 1
if self.n == self.N:
self.n = 0
Compare the results to the plain old Hit-and-Run step method: